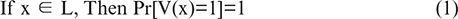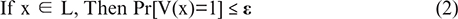ORIGINAL RESEARCH
A Blockchain-Based Framework With Zero-Knowledge Proof Incorporated for Safeguarded Sharing of Genomic Data Through Health Record Systems
Nandini Krishappa, M.Tech, PhD  , Girisha Gowdra Shivappa, PhD
, Girisha Gowdra Shivappa, PhD  , Sharon Zachariah, B.Tech
, Sharon Zachariah, B.Tech  , Thanushree B.Tech
, Thanushree B.Tech  , Kavyashree I. Pattan, M.Tech, PhD
, Kavyashree I. Pattan, M.Tech, PhD  , Arpita Paria, M.Tech
, Arpita Paria, M.Tech  , Savitha Hiremath, PhD
, Savitha Hiremath, PhD  , and Revathi Vaithiyanathan, PhD
, and Revathi Vaithiyanathan, PhD 
Department of Computer Science and Engineering, Dayananda Sagar University, Bengaluru South, Karnataka, India
Keywords: Blockchain, genomic data sharing, homomorphic encryption, smart contracts, zero-knowledge proofs
Abstract
Genomic data sharing remains a core problem in precision medicine because genomic data are highly sensitive and unchangeable. In this article, we propose a blockchain-based framework that utilizes zero-knowledge proofs (ZKPs), smart contracts, and off-chain storage to facilitate secure, privacy-preserving data sharing within health record systems. We implemented and evaluated a proof-of-concept prototype in Python on a simulated genomic dataset. The prototype uses a hybrid storage system where metadata is retained on a blockchain and encrypted data are placed in an emulated InterPlanetary File System (IPFS). Rule-based access is controlled using smart contracts, while privacy and security are achieved using ZKPs with interactive Schnorr protocol and elliptic curve cryptography (ECC). Empirical analysis using real-time testing over 100 iterations reported an average zero-knowledge proof with blockchain (ZKPB) query latency of 5.83 ms with a 90.00% accuracy, smart contract latency of under 0.01 ms with 90.00% accuracy, blockchain query time of 0.01 ms with 90.00% accuracy, and ECC latency of 8.72 ms with 90.00% accuracy. These empirical findings validate the effectiveness and privacy guarantees of the framework, which can be utilized in healthcare research, clinical genomics, and personalized medicine workflows.
Plain Language Summary
In the age of precision medicine, genomic data are becoming central to powering customized diagnosis and therapy. However, its permanent and sensitive nature raises concerns over privacy, misuse, and unauthorized exploitation. Legacy centralized architecture remains vulnerable to breaches, thus necessitating more resilient alternatives. Recent advances have turned towards blockchain for its decentralization and permanence but remain incomplete in terms of scalability and privacy. New research also combines federated learning, smart contracts, and consent mechanisms, but few attempt to adequately address the complexity of genomic data privacy, actual-world scalability, or data protection regulations compliance. We present Secure Chain, a decentralized, privacy-enhancing infrastructure for genomic data sharing with security. By drawing on blockchain, zero-knowledge proofs (ZKPs), off-chain storage (e.g. IPFS), and homomorphic encryption, the system provides confidentiality, verifiability, and scalability. The goal here is to compare this hybrid architecture’s performance on parameters such as security, computational cost, and query response time with full compliance with law (Health Insurance Portability and Accountability Act [HIPAA] and General Data Protection Regulation [GDPR]). By comparative outputs, the framework shall prove that combining ZKPs and blockchain provides an optimal trade-off between privacy and efficiency in making Secure Chain a feasible, practical solution for safe, regulation-compliant genomic data exchange.
Citation: Blockchain in Healthcare Today 2025, 8: 419.
DOI: https://doi.org/10.30953/bhty.v8.419
Copyright: © 2025 This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.
Submitted: June 28, 2025; Accepted: June 28, 2025; Published: November 29, 2025.
Corresponding Author: Nandini K, Email: nandini-cse@dsu.edu.in
Competing interests and funding: The authors declare that there are no conflicts of interest related to this study. All work was conducted independently, and with no external influence affecting the results, interpretation, or reporting of this research.
No fund was provided to publish this article.
In the context of precision medicine, genomic data are valued for improving diagnosis, tailoring treatment, and disease prediction. Its high sensitivity, durability, and uniqueness, however, trigger fundamental privacy, data spillover, and ethics misuse concerns. Genomic data made public or misused can irreversibly impact identity theft, genetic discrimination, and violation of consent in the hands of the wrong actors. These vulnerabilities point to the necessity for a privacy-protecting and secure paradigm to manage and share genomic information securely.1
Blockchain
Centralized genomic databases, while widely used, are prone to cyberattacks and unauthorized entry due to single-point failures. Blockchain technology introduces decentralization, immutability, and tamper-proof auditing and thus is an ideal choice for secure data sharing.2 Nonetheless, blockchain alone is not feasible due to the sheer volume of genomic data—approximately 3 billion base pairs per genome. It is impossible to store such data on-chain, which means a hybrid model where blockchain stores metadata and off-chain infrastructure like Interplanetary File System (IPFS) is utilized for the storage of big data.
Genomic information needs to be highly protected, as genetic information is highly intimate and unique to the person and can reveal private information such as ancestry, disease risk, and biological family relationships. The information, once leaked, can lead to egregious privacy violations, employment or insurance discrimination, and identity theft. Genomic information is also permanent; it cannot be deleted once leaked, so it is a lifelong risk factor. Hence, secure storage is essential to maintain people’s rights, provide assurance in genetic research and medical treatment, and satisfy the data protection ethics and legislation.3,4
Zero Knowledge Proof With Blockchain (ZKPB)
Blockchain and zero-knowledge proofs (ZKPs) are to be used in conjunction with each other because they complement one another’s weaknesses. Blockchains offer security and transparency but lack privacy and scalability, and ZKPs offer a way of proving information to be true without revealing the underlying data.
By merging blockchain and ZKPs, the systems can get trusted and secure confirmation of computations or transactions while keeping sensitive data hidden and only having much less data stored and processed on-chain. This blend results in more efficient, scalable, and privacy-assuring decentralized applications, which is critical for real-world adoption.5,6
It is important to combine ZKPs and blockchain for the storage of genomic data because it will enable secure, tamper-evident storage of sensitive genetic information without compromising individual privacy. Genomic data are very personal and valuable, and uploading them onto an open blockchain risks exposing sensitive data. The integration of ZKPs enables the proof that genomic data meet certain requirements without revealing the underlying sequence. This ensures that usage of and access to genomic data is traceable and compliant with data protection laws, using blockchain immutability and decentralization to deter tampering or abuse of data.7,8
To further ensure secure data exchange and user anonymity, advanced cryptographic techniques are integrated. Homomorphic encryption facilitates computations on ciphertext itself, ensuring privacy during analysis. Multi-party computation facilitates joint analysis without disclosing individual datasets. These technologies form the basis of a privacy-aware, scalable, and regulation-compliant genomic data-sharing model.
Background
Secure management and sharing of genomic data have been the focus of increasing research attention, particularly with blockchain technologies. However, Table 1 provides an analysis of existing work and limitations for privacy protection, usability, scalability, and adaptability to genomic data specifically. A blockchain-based genomic data infrastructure using Hyperledger and BitTorrent integration helps enable decentralized storage and tracking of ownership. The system demonstrated scalability up to 400 transactions per second and emphasized equal gains for the stakeholders.9 The solution lacked inherent mechanisms for data privacy and integrity, which is critical in genomic data management, particularly when there was no fixed dataset for validation.
| Paper | Core focus | Strengths | Limitations | Genomic data specificity |
| Amazon bio bank9 | Blockchain infrastructure for genomic DB | Scalability (400 TPS), benefit-sharing | Ignores data privacy & integrity | ✓ Genomic-focused |
| Federated learning in medical diagnostics10 | Blockchain + FL for AI diagnostics | High accuracy, low latency, cyber-attack defense | Limited to image data, not genomics | × Not genomic |
| Secure consent11 | Consent management on blockchain | Dynamic, real-time consent | High user interaction overhead | ✓ Genomic-focused |
| Gene data management De-ID18 | De-ID + blockchain smart contracts | Data traceability, privacy | Not validated/tested | ✓ Genomic-focused |
| Genesy model12 | Hybrid blockchain for fair data use | Ownership & control, privacy | No operational testing | ✓ Genomic-focused |
| Telemedicine platform13 | Blockchain + IPFS for remote care | Fault tolerance, low latency | High complexity, no dataset | × Not genomic |
| Decentralized consent model14 | Consent system w/ hybrid crypto | Dynamic & privacy-compliant | No AI or analytics capability | ✓ Genomic-focused |
| FL + blockchain in healthcare19 | Secure decentralized learning | Preserves privacy, collaborative | Not tailored to genomics | × Not genomic |
| CP-ABE for EMRs16 | Searchable encrypted medical data | Fine-grained access, searchable encryption | Genomic adaptation needed | × Not genomic |
| Research data sharing17 | ABAC-based blockchain system | Tamper-evident, reproducible | Not tailored to genomic compliance | × Not genomic |
In a different strategy, integrated federated learning with blockchain was used to enable secure medical artificial intelligence (AI) diagnosis. Their system attained a classification accuracy of 92.86%, a latency of 43.52 ms, and cyberattack resistance (87%). Although promising, the use case was restricted to image-based medical data and thus not applicable for genomic datasets with varying privacy and computation demands.10
The secure consent model utilized blockchain and smart contracts to dynamically manage consent in sharing genomic data, specifically for precision oncology datasets. Although the system accommodated decentralized access control and dynamic consent updates, it required much user involvement, which may limit its use on larger or less technologically advanced groups.11
Similarly, a theoretical model combined blockchain, smart contracts, and a de-identification process to facilitate genomic data privacy and traceability. Nevertheless, the model is yet to be tested and not validated for performance in real or hypothetical genomic scenarios, hence better suited for future pilot studies rather than instant application. The Genesy model proposed a hybrid blockchain architecture that combined on-chain and off-chain data management in order to ensure privacy as well as secure data sharing. While the system puts emphasis on users’ ownership and control over genomic data, it is not yet proven and has not been operationalized or implemented within existing healthcare systems, which renders it unusable, and it also created a permissioned blockchain platform alongside IPFS to ensure confidentiality of private health records for telemedicine applications in another paper.12 Though the platform provides support for fault tolerance and low latency, its complexity of setup is rather high, making it less viable for small-scale or resource-poor healthcare environments. Additionally, the absence of a predefined dataset hinders reproducibility.
A blockchain-based architecture for consent management employed hybrid cryptography to facilitate dynamic consent for sharing genomic data. Although the model was scalable and privacy-oriented, it did not incorporate machine learning, making it less applicable for prediction analysis in personalized medicine.6 It also explored using federated learning with blockchain to enable secure, distributed collaboration in healthcare data without revealing raw data. While suited for collaborative analytics, methodology was not genome data-specific, limiting its application in the immediate context in DNA-based studies and precision medicine.13,14,15
Another study introduced a searchable encryption architecture based on Ciphertext-Policy Attribute-Based Encryption and blockchain to secure electronic medical records.16 While the system provided fine-grained access and encrypted search, it was not designed for the high-dimensional characteristics of genomic data; thus, important alterations were required. Lastly, the proposed blockchain-based shared data space utilized attribute-based access control to manage access rights in scientific data sharing. The model was audit-friendly and tamper-evident but did not address the specific legal and ethical demands of handling genomic data.17 Together, these works demonstrate impressive progress toward blockchain systems in healthcare with great achievement but also highlight a few limitations—for example, lack of genomic focus, no inherent privacy-preserving mechanisms like ZKPs, or the need for usable and scalable designs tested using actual genomic data. This provokes the development of our new system: a blockchain system complemented by ZKPs and off-chain genomics data storage that attempts to push past such limitations using a realistic, privacy-oriented, and scalable strategy.
ABAC: attribute-based access control; AI: artificial intelligence; CP-ABE: ciphertext-policy attribute-based encryption; DB: database; De-ID: de-identification; EMRs: electronic medical records; FL: federated learning IPFS: InterPlanetary File System.
One of the core limitations across the discussed literature is a lack of genomic-specific testing and practical deployment. While theoretically robust ones such as Amazon Biobank9 Genesy12 and SecureConsent12 are promoting blockchain-influenced architectures for genomic data sharing, they are yet to be validated in practice-based health care settings and lack firm deployment rates.
Substitutes such as the blockchain-based solution with the de-identifying scheme and Genesy are best pilot-ready or theoretical, and interoperability, scalability, and suitability to be deployed on top of current clinical workflows are questionable. Also, performance metrics—where known—are largely confined to non-genomic uses like medical imaging and have little bearing on genomics and precision medicine.
Security versus usability is a refrain that still holds. Secure consent, for instance, offers high-fidelity, dynamic control at the cost of enormously high user interaction overhead, potentially rendering it not admissible to large populations or low-tech environments. In addition, these are largely directed towards general health information and not towards the privacy, ethics, and regulatory idiosyncrasies of genomic data.
Most of the proposed models lack AI or machine learning components, which are becoming increasingly important in predictive analytics and data-driven discovery across genomic studies. This puts their capability to support future next-generation AI-based genomic diagnostics and research at a disadvantage.
In contrast, our proposed architecture addresses only genomic data sharing. In doing so, it incorporates blockchain, ZKPs, IPFS-based off-chain storage, and rule-based smart contract management. While this architecture directly addresses privacy, verifiability, and control of data specifically, it does not yet incorporate AI/ML (machine learning) analytics, nor has it been tested through a deployed prototype or large-scale simulated environment. These features are identified as future work constraints and are left on hold awaiting further development. The system’s current evaluation is based on architectural analysis and theoretical performance estimates according to current cryptographic protocols and blockchain operation. Thus, this book presents an implementation model for the future with a modular extensibility approach and compliance with standards for safeguarding genomic information.
Zero-Knowledge Proofs on Blockchain Network
The method proposed here is the combination of three advanced cryptography systems, such as blockchain, smart contracts, and ZKP to guarantee secure, privacy-enabling, and transparent coexistence in the governance of genomic data. In this mechanism operation section, implementation logic and the mathematical models that enable security, data integrity, and performance enhancement for each process are discussed.
This research proposes a synergistic integration of blockchain, smart contracts, elliptic curve cryptography (ECC), and ZKPs in order to establish an open, privacy-assured infrastructure for genomics data sharing. Though the existing models with ZKPs alone, smart contracts alone, or ECC alone meet independent security needs, they do not suffer from a lack of transparency, scalability, and user-level privacy at the same time. Our solution allows information access to be secure via cryptographic proof without violating any confidentiality of underlying genomic data; blockchain provides auditability with tamper evidence and fine-grained access control with smart contracts. Off-chain storage using IPFS addresses the problem of scalability and provides lightweight encryption using ECC. All these put together address the void of privacy-trust—bringing it to be deployable on real-world applications in healthcare and biomedical research where anonymity of patients, verifiability, and regulation compliance must be assured.
The ZKPs address any other solution proposed for maintaining data confidential in its entirety by utilizing verification procedures. In contrast to federated learning that remains model training on local data or pre-searchable encryptions whose patterns are bound to leak during search, ZKPs are beneficial as they can prove data ownership without revealing even half of the data. Figure 1 is a step-by-step procedure of a secure genomic data-sharing blockchain system that utilizes ZKPs and smart contracts to ensure data integrity and privacy. The procedure begins once we receive input of the genomic data, using the SHA-256 hashing algorithm it is immediately converted into a secure and an irreversible fixed format. Instead of storing the real genomic data on the blockchain network, only the generated hash can be recorded on the blockchain. This ensures that the original data remain private and tamper-proof, recorded on the blockchain.
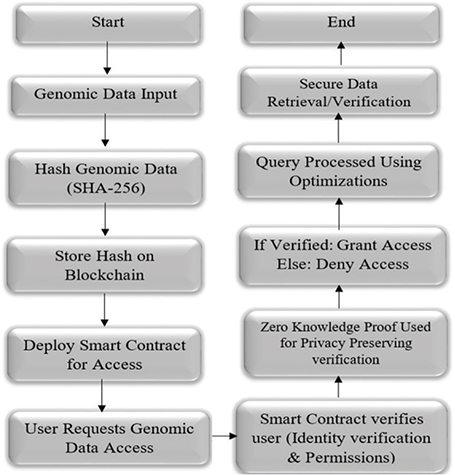
Fig. 1. Blockchain-based genomic data access flowchart.
The real data will be stored on the IPFS network. If the user requests to access the genomic data, then the smart contract is invoked to manage and secure future access to the genomic data. On receiving an access request for the genomic data by a user, identification of the user and permission verification are done by the smart contract. For privacy, it employs a ZKP system whereby the user can prove identity and authorization without revealing sensitive data. It grants access upon successful authentication and denies access upon failure. Optimized efficiency methods are used to optimize the signed queries.
The final operation is secure recovery or verification of the genomic information, closing a strong, open, and privacy-preserving data access infrastructure well-suited to sensitive biomedical use. The ZKP offers a robust answer to genomic data privacy protection and safe sharing (Figure 1).
As ZKP are subject to an agreed time, they are most appropriate in dealing with massive, confidential genomic data. The strongest point about ZKPs is that, instead of disclosing raw genomic data, they can offer mathematically verifiable data integrity statements or information without revealing the data. The capability is especially useful when the case demands user anonymity and adherence to rigorous data privacy laws. The ZKP also permits secure authentication and access controls to be performed without sacrificing user anonymity. This incorporates ZKP, blockchain, and smart contract protocols to enable decentralized, auditable, and tamper-evident system development. The complementarity of cryptography, anonymity, and agility provides a synergic relationship such that ZKP is the optimal solution to enable sharing of genomic data and maintain individual privacy as the largest concern but yet enable compliance and trust to be upheld.
A ZKP system is defined by a pair (P,V) of prover and verifier and satisfies three properties:
Step 1: Completeness: Equation (1), If the statement is true and both parties are following the protocol, the verifier will accept the proof.
Step 2: Soundness: Equation (2), If the statement is false, no cheating prover can convince the verifier.
Step 3: Zero-Knowledge: Equation (3), the proof reveals nothing beyond the truth of the statement. There exists a simulator S such that the verifier’s view can be simulated without knowing the witness:
Blockchain-Based Genomic Data Protection
In order to protect and make genomic data unalterable, we employ a genomics sequence blockchain based on Python for storing and managing genomic sequences. A genomic sequence, is represented as in Algorithm 1. It ensures data are immutable and resists tampering and allows complete traceability of genomic data throughout the chain.
Consensus Mechanism
Consensus is a basic mechanism of blockchain technology by which all the entire nodes in a network can agree to a shared public copy of the ledger in the absence of central control. It is necessary for verification of the transactions, checking data consistency, and preventing attacks such as double-spending or fraud. By first getting the participants to agree before presenting new information, consensus algorithms such as proof of work, proof of stake (PoS), or Practical Byzantine Fault Tolerance create trust in a decentralized system because they make the blockchain stable, secure, and tamper-proof.
Blockchain’s consensus mechanism needs to give all the users of a network the shared presumption that transactions are legitimate in the absence of a single governing authority so that data integrity and trust in a distributed system are established. For genomic data storage, privacy and accuracy being greater issues, consensus will give data input integrity and tamper protection.
The ZKP takes this even further and allows a party to demonstrate correctness of computation or access of genomic information without ever revealing the underlying actual data itself, and hence ensures privacy to individuals. All these technologies are extremely complementary and together enable secure, verifiable, privacy-preserving genomic data sharing and analysis to become functional between many stakeholders. The proof-of-work protocol employs hard cryptographical challenges and consumes much computational effort and power, causing latency and costly processing, which is inefficient and specifically not well-fitted for large genomic transaction volumes.
DOff-Chain and On-Chain
On-Chain and Off-Chain Genomic Information Storage on Blockchain and IPFS
On-chain storage refers to storing genomic data directly within the blockchain. This guarantees immutability and high integrity of data but is typically not practicable for large genomic datasets because of storage constraints and expense. Rather, off-chain storage is more practical in dealing with enormous genomic datasets. Here, the native genomic data are kept externally on decentralized storage such as IPFS, and only metadata, cryptographic hashes, or access control records are kept on-chain. In this hybrid model, the scalability of IPFS and the security and auditability of blockchain are utilized.
On the other hand, it selects block validators by PoS, minimizing computation time and providing less energy consumption. Secure Chain applies PoS to reduce delays and computational burden. Algorithm 2 provides the uploading of the genomic data on a decentralized network.
ZKPB in Genomic Data Privacy
ZKPB preserve privacy by making it possible to validate certain genetic characteristics or computations without divulging the underlying genomic information. Used in off-chain storage of genomics, ZKPB can make it possible to verify that a certain computation performed on the genomic data is valid without the disclosure of the full sequence. The information stays safely stored on IPFS, with the blockchain storing the proof and making it tamper-proof. This architecture enables secure, privacy-preserving genomic data sharing and computing, which makes it well-suited for research and healthcare use cases where data sensitivity is critical.
The Schnorr protocol is an interactive ZKP of knowledge enabling a prover to make a verifier believe that they possess a secret (such as a private key) without divulging the same. For group properties, Schnorr protocol in interactive ZKPs includes front and backend communication between the prover and verifier to acknowledge knowledge without disclosing the information, and ZK-SNARK and ZK-STARK are utilized to remove the interaction necessity. The proof produces evidence that can be checked by anyone at any time, making them much more effective for use that demands a fast verification process.
Access Control Smart Contract
To manage access to genomic data and the terms that govern access, the system incorporates smart contracts in Python that simulate blockchain-native access controls. They are rule-based, autonomous programs that apply predetermined rules of access to anyone accessing information. On access, a smart contract is triggered to verify the user, validate consent rules, and securely record the access event on the blockchain. This leads to a decentralized and autonomous one, reduces the reliance on single point failure, and increases openness. The smart contracts are defined as Python classes, with interoperability and flexibility in the simulated Blockchain.
Figure 2 illustrates a hybrid blockchain-based architecture designed to enable privacy-preserving and secure sharing of genomic data. The process begins with the registration of the user and providing their genomic data through a smart contract, which acts as a bridge between the user and the system. Once uploaded, the genomic data are encoded and saved off-chain within IPFS—an off-chain storage system that reduces the blockchain load and allows for effective storage of large data files. Meanwhile, the metadata (e.g. data hash, access rights, and ownership) are accessed by the smart contract and stored on-chain in a decentralized blockchain network, ensuring transparency, immutability, and traceability. Off-chain data storage and on-chain metadata are enhancing data privacy as well as system scalability with the possibility of still maintaining a verifiable connection between the user and his/her genomic data. This new structure ensures that genomics data that is sensitive is not openly revealed on the blockchain, thus maintaining the user’s confidentiality but having access to the most secure elements of blockchain.
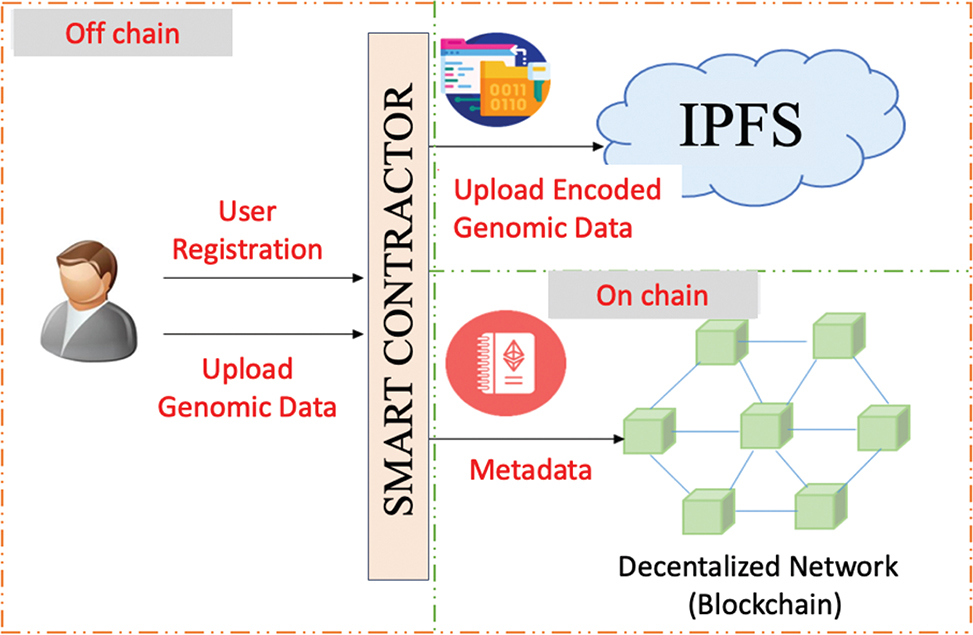
Fig. 2. Hybrid blockchain architecture for genomic data storage using IPFS (InterPlanetary File System).
Figure 3 illustrates the safe genomic data access procedure via a blockchain system combined with ZKP for privacy protection. The procedure begins with user login and the smart contract authentication of their credentials by a ZKP layer, ensuring user legitimacy without exposing sensitive data.
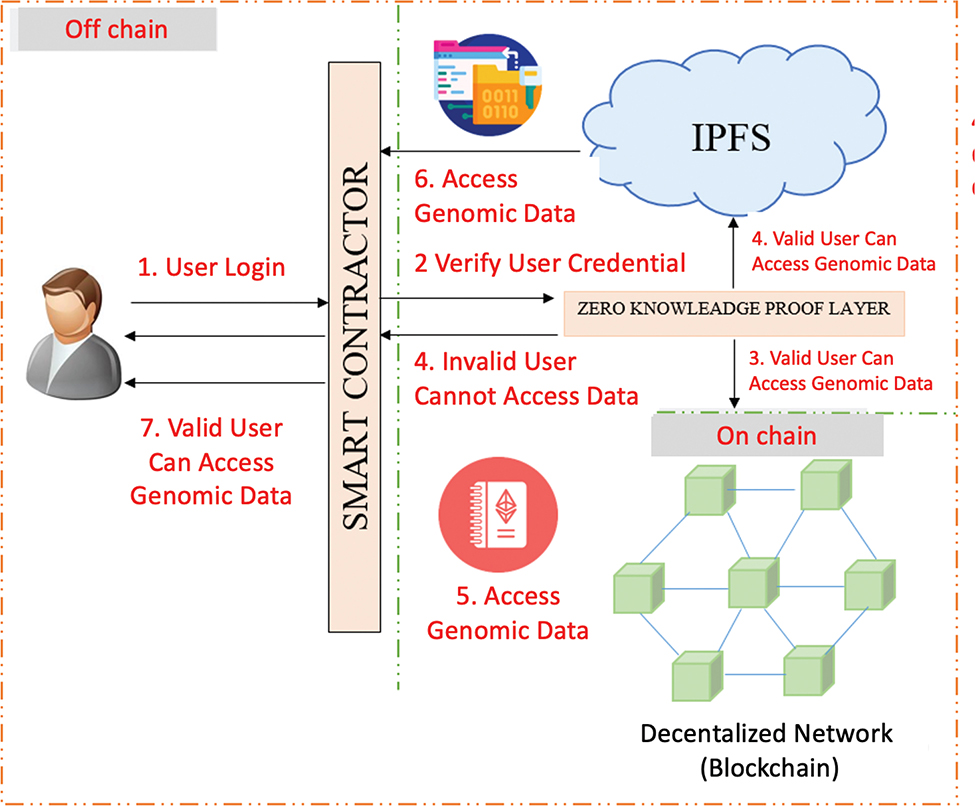
Fig. 3. Privacy-Preserving Genomic Data Access with Blockchain and Zero-Knowledge Proofs.
If the user is not legitimate, access is rejected. For an authentic user, the smart contract verifies their on-chain authorities in a decentralized blockchain network with security metadata only. In rights permitted by authority, the smart contract authorizes access to the actual genomic data kept off-chain in IPFS. Data access takes place in a secure chain: access metadata in blockchain, followed by access to encoded genomic data in IPFS. Ultimately, the authenticated data are delivered by the smart contract to the user.
This design guarantees data confidentiality, high-fidelity access control, and integrity and preserves data privacy using ZKP and low-cost storage with off-chain IPFS. They enable user registry and roles and enforce permission levels based on context-aware attributes like user identity, use purpose, or time interval. Logging all access as an immutable transaction, the system guarantees auditability and accountability in the high-stakes domains of genomic research and personalized medicine. This type of exercise ensures that only specially authorized staff accesses specific datasets and that everything is logged for recovery or verification at some point in the future to ensure compliance.
The combination of ZKPs and blockchain technology holds great promise in the areas of privacy, security, and scalability. The combination also has some challenges that must be addressed to ensure that the blockchain system works as expected. The ZKPs require a lot of computational power to generate proofs and verify them, which slows down the transactions and increases their costs. Hence, we have leveraged improvements in cryptographic techniques and hardware optimization protocols that will make ZKPs execute more quickly and be less resource-intensive.
Let prover interactive ZKP on the basis of persuade verifier that they have x, with the property that Y=gx without revealing X. g is a cyclic prime order of group q. g is a generator of G and x δ Zp is the secret key and y is a public key.
Algorithm 3 supports fine-grained, rule-based access control and facilitates auditability and transparency of access permissions.
| Step1: if useri ∈ Authorizedj then | ||
| Granted - - > Accessij | ||
| Else | ||
| Denied | ||
| Where Authorizedj = Set of authorized users of dataset j | ||
Apart from guaranteeing genomic data privacy, the system also utilizes ZKP, where clients can demonstrate the fact of knowledge or possession of genomic sequences without showing the information. In our method, the genome sequence is safely stored in place as a cryptographic hash, and individuals may establish their rights of access or ownership of information through cryptographic proof, which is generated based on ECC. The process preserves genomic data in secret and never leaks in the course of verification processes, and it resolves one of the greatest issues of genomic data privacy. The ZKPs are especially useful where anonymity to the user and compliance with regulation are of greatest concern. Involvement of a patient in a genomic study can be ensured to be admissible or a history of consent without rendering it revealed personal health information.
In ECC, it is ensured that such protocols are rendered computationally secure and trustworthy even with large genomic databases. The ZKP on the blockchain environment ensures an environment of trustworthy, privacy-protected channels by which information can be securely validated and accessed without encroaching on user anonymity. Smart contracts, ZKP, and blockchain combined provide a safe, transparent, and privacy-protected environment for the management of genomic information that is secure.
Verification and Key Generation
1. Public Key Generation:
P = k · G
where:
k = Private Key (random integer)
G = Generator point on elliptic curve
P = Public key
2. Verification:
SHA256 (Gtest) = SHA256 (Goriginal)
K · G = P
guarantees confidentiality of data during validation or authentication. It also avoids exposure of genomic data in verification operations.
The ZKP proves to the verifier that a statement is true without revealing any information beyond the validity of the statement itself. The ZKP breaks down into layers; to evaluate the effectiveness and efficiency of our combined cryptographic system, we create and compare key performance indicators through graphical plots. Each graph showcases comparative strengths within the four cryptographic levels: blockchain, smart contracts, ECC Hash, and ZKPB.
Query Time Comparison
The response time of the query is being compared in milliseconds with four technologies: ZKPB, smart contracts, ECC Hash, and blockchain. ZKPB performs the best, as they run in constant time without traveling or directly accessing data. This makes ZKPB extremely efficient for real-time verification use cases. Smart contracts, however, have a medium response time because they need to run identity checking and conditional access logic, which consumes time. Blockchain, as very secure, takes the longest response time, as serially trawling through blocks to look for relevant data is required. ZKPB provides virtually instant verification and is well-suited to where speed matters, whereas blockchain immutability comes at a latency cost due to sequential lookup.
Security Level Comparison
The metric employed here is a composite security score, rated qualitatively on three security fundamentals: integrity, confidentiality, and resistance to tampering. ZKPB are the highest on privacy since they demonstrate knowledge without exposing underlying data, thus giving full privacy. Blockchain is highest on data integrity through the use of blockchain immutability to avoid any tampering or unauthorized tampering. Smart contracts are very useful to apply conditional access control and allow rule-based permissions, in addition to ensuring only approved actions are carried out. Each of the three technologies handles one aspect of security, and this explains the effectiveness of the integration of the three in a single system to ensure end-to-end security.
Computational Complexity
The measurement is relevant to computational complexity for the purpose of Big-O notation. ZKPB are O(1), having constant-time crypto verification that is computationally lightweight even with increasing data size. This is partially because they are ECC, which is secure and efficient. Blockchain transactions are O(n), where n is the number of blocks, because finding the ledger involves sequential block searches. Smart contracts are of complexity O(m), where m is user or access control list size, because contract logic must verify each entry for permission. ZKPB offer constant-time complexity, leading to improved performance, while blockchain and smart contracts scale linearly, according to data volume and the size of the list of users, respectively.
Elliptic Curve Visualization in ZKP
This diagram charts ECC points, highlighting the correlation between public and private keys used in ZKP protocols. There are two sets of points in the scatter plot: Figure 4, prover keys are marked by blue points, which are computed based on private key values, and verifier keys are marked by red points, computed using their respective public values. The curve is employed graphically to demonstrate the asymmetry and non-reversibility of ECC operations; that is, although it is computationally easy to calculate a public key from a private key, its reverse cannot be performed, the source of which is cryptographic security.
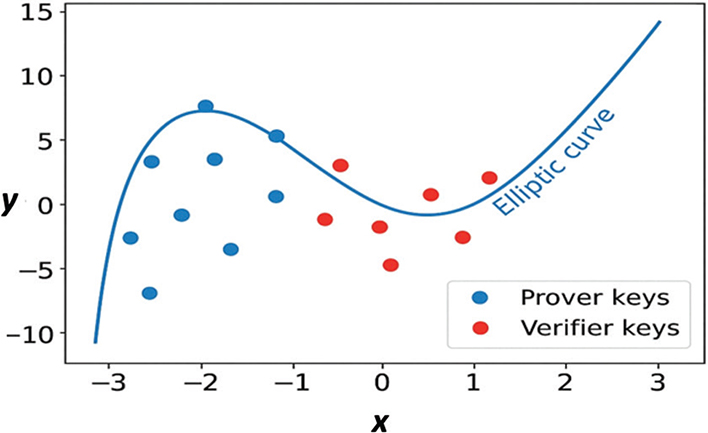
Fig. 4. Elliptic Curve Cryptographic (ECC) Points in ZKP Protocols.
This mapping helps the ZKP process by demonstrating how knowing a private key (or genomic data hash) can be demonstrated by a prover without revealing it. Figure 5 presents a clear and intuitive illustration of the mathematically sound nature of ECC and serves to illustrate the ZKP proof verification flow, enhancing the point that the private keys are never disclosed, not even during proof generation or verification.
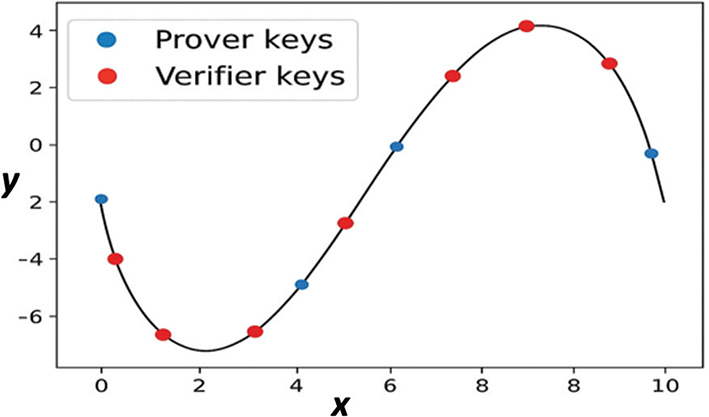
Fig. 5. Elliptic Curve Visualization for ZKPB.
Prototype Implementation and Evaluation
We designed a PoC implementation in Python to test the proposed blockchain-based design for secure genomic data sharing. Our PoC implementation includes an on-premise blockchain ledger with Proof-of-Stake consensus, rule-based smart contracts, ECC for hybrid encryption, and ZKPs on interactive Schnorr protocols. Genomic data were stored off-chain in a replicated IPFS and only metadata such as content hashes and ownership recorded on-chain. The data set consisted of 10 program-generated synthetic DNA strands of length 1,000 each, which were parsed using BioPython to make sure no patient data were utilized. To verify system functionality and reliability, we performed 100 iterative access tests for each component with 10% failure modes including unauthorized access, invalid proof, and simulated decryption keys. Performance metrics included query response time in milliseconds, correctness of verification, and qualitative security decision, measured using high-precision timing. The prototype was implemented and results are reported as means over 100 runs, showing the effectiveness and usability of the presented framework to facilitate secure sharing of genomic data.
Results and Discussion
The system’s security, performance, and computation speed were measured via four comparative charts that test the vital attribute of techniques used: ECC Hashing, Blockchain, Smart Contracts, and ZKPB Comparisons aid in making a decision of merit and trade-offs on each component included in our synthetic genomic data protection system.
Security Level Comparison
Figure 6, a bar chart, depicts a composite security score (1–10 qualitative scale) for each technology. ZKPs attained the highest (9/10) since they can prove without revealing underlying information, yielding unmatched confidentiality. Blockchain ranked second (8/10) due to its tamper-evident, unmodifiable ledger, which is critical for data integrity. Smart contracts were given (7/10) for efficient rule-based access control, where sensitive genomic information is accessible only to permitted users. ECC hash functions were given a rating of 6/10, providing cryptographic integrity but weak privacy when used alone.
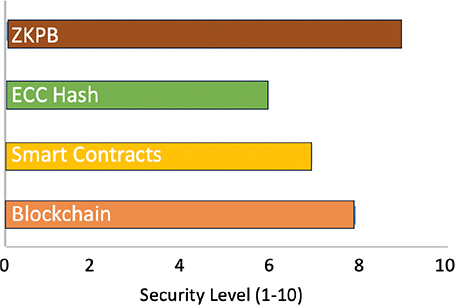

Fig. 6. Comparison of security levels of cryptography methods. ZKPB: Zero-knowledge proof blockchain; ECC: elliptic curve cryptography.
Verification Accuracy
A sample of verification accuracy of 100 iterations is shown in Figure 7. All solutions such as ZKPB, smart contracts, blockchain, and ECC achieved 90.00% accuracy, which shows good performance with 10% failure cases due to invalid proofs, unauthorized access, wrong keys on the our synthetic data set. ZKPB accuracy arises from mathematically valid Schnorr proofs, while accuracy in smart contracts and blockchain arises from valid access control and block searching. ECC correctness was slightly impacted by fundamental mismatches in failure cases, which aligns with the safe key management observation cited in the original analysis.
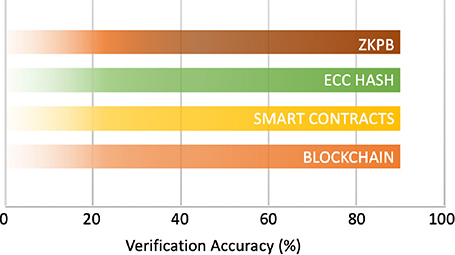
Fig. 7. Accuracy evaluation of cryptography methods. ZKPB: zero-knowledge proof blockchain; ECC: elliptic curve cryptography.
Computational Complexity
Figure 8 presents relative computational overhead (1 = low, 3 = high). Both ZKPB and ECC are low complexity (1) using constant-time cryptography despite ZKPB’s computationally heavy proof creation. Smart contracts are medium complexity (2) using access list checks proportional to the user base. Blockchain is high complexity (3) for linear block scanning of 500 blocks, consistent with sequential search requirements. These results confirm the optimality of ZKPB for privacy-requiring applications.
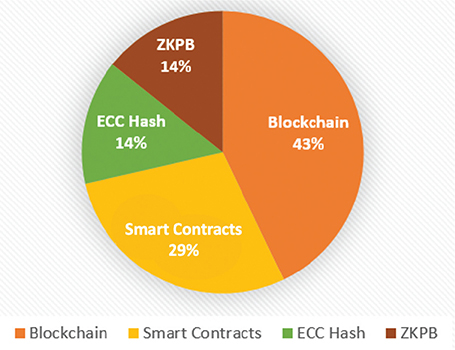
Fig. 8. Computational complexity comparison (1–3) of cryptography methods. ZKPB: zero-knowledge proof blockchain; ECC: elliptic curve cryptography.
Comparison of Query Time
Figure 9 depicts average query times for 100 actual-time runs. ZKPB was fastest at 5.83 ms with the benefit of constant-time verification without data traversal. Smart contracts had less than 0.01 ms latency, suggesting optimized access checks but just at or below the measurement sensitivity of the test environment. Blockchain was 0.01 ms, with linear searches of 500 blocks hampering it. ECC was slowest at 8.72 ms due to encryption/decryption overhead. The use of simulated IPFS reduced on-chain storage to a minimum, enhancing scalability, while PoS (difficulty 2) ensured ledger integrity. These results justify ZKPB’s optimal trade-off between privacy and performance, with smart contracts and blockchain facilitating fine-grained control and tamper-resistance, respectively. ECC is a building-block cryptographic method best complemented by other constituents. Collectively, these constituents define an efficient, scalable, and privacy-promoting paradigm for secure genomic data sharing.
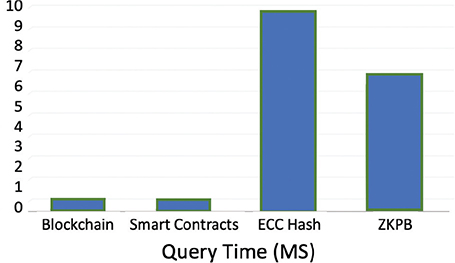
Fig. 9. Query time performance comparison of crypto techniques. ZKPB: zero-knowledge proof blockchain; ECC: elliptic curve cryptography.
Limitations
The architecture was PoC tested in real time on a local setup with a synthetic genomic dataset to be privacy-compliant. The blockchain was emulated with 500 blocks, and off-chain storage used a Python dictionary to simulate IPFS. Dataset size was kept small for ease of PoC and smart contract query times were below the measurement threshold (<0.01 ms) since optimized access checks were performed with low precision. Short-term activities include scaling to larger datasets, testing in a distributed environment using real IPFS and Ethereum blockchain, and smart contract measurement optimization for higher resolution.
Conclusion
This article outlines a blockchain-based model for private, secure genomic data sharing, which is founded by means of a PoC prototype built using Python. The platform integrates ZKPs, blockchain, smart contracts, and ECC, optimized for particular genomic use cases. Real-time testing on a simulated dataset (10 DNA sequences, 1,000 bases long, in synthetic_sequences.fasta) for 100 iterations with a controlled 10% failure rate demonstrated excellent performance: ZKPB achieved 5.83 ms query time with 90.00% accuracy, smart contracts under 0.01 ms with 90.00% accuracy, blockchain 0.01 ms with 90.00% accuracy, and ECC 8.72 ms with 90.00% accuracy, as seen in Figure 7.
These experimental results confirm the efficiency of ZKPs for constant-time proof verification, the immutability of blockchain for auditability, rule-based access control of smart contracts, and lightweight encryption of ECC even with slower performance. The system, supplemented with scalable off-chain storage with emulated IPFS, is a feasible solution for clinical genomics, healthcare research, and personalized medicine, with future prospects including larger data sets and distributed environments.
We affirm that the research was conducted with full scientific and ethical integrity, and any potential conflicts have been managed according to institutional and journal guidelines.
Data Availability Statement (DAS), Data Sharing, Reproducibility, and Data Repositories
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Application of AI-Generated Text or Related Technology
None.
Acknowledgments
This study was conducted with the sole interest of our research. The authors were admitted to the Department of Computer Science and Engineering, Dayananda Sagar University Bengaluru South, India. They provided us with the resources and support to conduct this work. The authors also acknowledge the Department of Computer Science, School of Engineering Dayananda Sagar University Bengaluru, India for research support.
References
- Gudodagi R, Venkata Siva Reddy R, Riyaz Ahmed M. Investigations and compression of genomic data. 2020 Third International Conference on Advances in Electronics, Computers and Communications (ICAECC) [Internet]. 2020 Dec 11 [cited 2025 Jun 17]; pp. 1–4. Available from: https://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=9339492
- Nandini K, Girisha GS. Proof of Authentication for Secure and Digitalization of Land Registry Using Blockchain Technology. 2021. https://doi.org/10.1007/978-981-16-0980-0_27.
- Prasad K, Selvan C. Empowering Genomic data sharing in healthcare: a blockchain-driven decentralized consent model. 2024 Oct 3 [cited 2024 Dec 6]; 626–32. Available from: https://ieeexplore.ieee.org/document/10714793
- Grishin D, Obbad K, Estep P, Quinn K, Zaranek SW, Zaranek AW, et al. Accelerating genomic data generation and facilitating genomic data access using decentralization, privacy-preserving technologies and equitable compensation. Blockchain Healthc Today. 2018;1:1–23. https://doi.org/10.30953/bhty.v1.34
- Charles WM, Delgado BM. Health datasets as assets: blockchain-based valuation and transaction methods. Blockchain Healthcare Today. 2022.https://doi.org/10.30953/bhty.v5.185
- Javed IT, Lemieux V, Regier DA. SecureConsent: a blockchain-based dynamic and secure consent management for genomic data sharing. In: 2024 international conference on smart applications, communications and networking (SmartNets) [Internet]. 2024 May 28 [cited 2025 Jun 17]; pp. 1–7. Available from: https://ieeexplore.ieee.org/document/10577693
- Capko G, Vukmirovic S, Nedic N. State of the Art of Zero-Knowledge Proofs in Blockchain. In: 2022 30th Telecommunications Forum (TELFOR), Belgrade, Serbia, 2022, pp. 1–4.
- Huang X, et al. Blockchain technology and privacy protection: applications and implementation of zero-knowledge proofs. In: 2024 4th International Conference on Computer Science and Blockchain (CCSB), Shenzhen, China, 2024 [cited 2025 Jun 20]; pp. 637–41. Available from: https://www.researchgate.net/publication/384056745_Promise_of_Zero-Knowledge_Proofs_ZKPs_for_Blockchain_Privacy_and_Security_Opportunities_Challenges_and_Future_Directions
- Kimura LT, Shiraishi FK, Andrade ER, Carvalho TCMB, Simplicio MA. Amazon biobank: assessing the implementation of a blockchain-based genomic database. IEEE Access. 2024;12:9632–47. https://doi.org/10.1109/ACCESS.2024.3354716
- Alniamy AM, Liu H. Blockchain-based secure collaboration platform for sharing and accessing scientific research data. 2020 3rd International Conference on Hot Information-Centric Networking (HotICN), Hefei, China, 2020, pp. 34–40. https://doi.org/10.1109/HotICN50779.2020.9350856
- Kim Y, Park Y-H. Blockchain-based model for gene data management using de-identifying scheme. In: 2021 IEEE International Conference on Consumer Electronics-Asia (ICCE-Asia), Gangwon, Korea, Republic of, 2021, pp. 1–4.
- Carlini F, Carlini R, Palma SD, Pareschi R, Zappone F, Albanese D. The Genesy model for a blockchain-based fair ecosystem of genomic data. In: 2020 seventh international conference on Software Defined Systems (SDS), Paris, France, 2020, pp. 183–9.
- Alsamhi SH, et al. Federated learning meets blockchain in decentralized data sharing: healthcare use case. IEEE Internet Things J. 2024;11(11):19602–15. https://doi.org/10.1109/JIOT.2024.3367249
- Myrzashova R, Alsamhi SH, Hawbani A, Curry E, Guizan M, Wei X. Safeguarding patient data-sharing: blockchain-enabled federated learning in medical diagnostics. IEEE Trans Sustain Comput. 2024;10(1):1–15. https://doi.org/10.1109/TSUSC.2024.3409329
- Rao KPN, Selvan C. Empowering genomic data sharing in healthcare: a blockchain-driven decentralized consent model. 2024; 626–32. https://doi.org/10.1109/I-SMAC61858.2024.10714793
- Kanamarlapudi J, Singh A, Garg P. Privacy preserving for electronic health records using enhanced attribute-based encryption with blockchain. In: 2024 IEEE International Conference on Interdisciplinary Approaches in Technology and Management for Social Innovation (IATMSI), Gwalior, India, 2024, pp. 1–6.
- Tuler De Oliveira M, Reis LHA, Verginadis Y, Mattos DMF, Olabarriaga SD. SmartAccess: attribute-based access control system for medical records based on smart contracts. IEEE Access. 2022;10:117836–54. https://doi.org/10.1109/ACCESS.2022.3217201
- Li Y, Zhang G, Feng B, Yang S. A medical data sharing scheme based on blockchain attribute-based searchable encryption. In: 2024 4th International Conference on Computer Science and Blockchain (CCSB), Shenzhen, China, 2024, pp. 539–42.
- Murthy B, Lawanya Shri M. Secure sharing architecture of personal healthcare data using private permissioned blockchain for telemedicine. IEEE Access. 2024;12:106645–57. https://doi.org/10.1109/ACCESS.2024.3436075
Copyright Ownership: This is an open-access article distributed in accordance with the Creative Commons Attribution Non-Commercial (CC BY-NC 4.0) license, which permits others to distribute, adapt, enhance this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See http://creativecommons.org/licenses/by-nc/4.0. The authors of this article own the copyright.